Editor’s Note – This essay was subsequently published as: Dillon, R.T., Jr. (2023b) Startled by Fontigens, sort-of, I suppose. Pp 217 – 223 in The Freshwater Gastropods of North America Volume 6, Yankees at The Gap, and Other Essays. FWGNA Project, Charleston, SC.
It takes a lot of forbearance to follow the FWGNA Blog. I am aware of this. I break every rule of social media – exploring arcane topics at great depth, extending the attention span of my readership over many months. Sometimes years. It is not uncommon for me, your petulant host, to expect you to remember some petty anecdote or obscure factoid that may have flickered through my brain years ago, and if you do not remember it, scold you and assign it for homework. And then forget it, myself.
So, welcome to snail science – a gradual unfolding, a cautious move forward, a startling, and an extended period of dormancy.
The stimulus for our gradual unfolding this month is the publication late last year of a paper by Liu, Schroeder, Berry, and Dillon on sequence divergence in phreatic hydrobioid snails of the genus Fontigens [1]. You might be forgiven if your memory of my 2006 essay on the Springsnails of the Blue Ridge is a bit fuzzy. But I do trust that you remember our three-part series on Lori Schroeder’s tiny snails back in 2017, yes? And surely, our July 2019 report of the discovery of a single putative Fontigens cryptica in a spring at the Bernheim Forest in central Kentucky is fresh on your mind. And you must have read my Bob Hershler tribute last month, right? Don’t tell me that you didn’t.
So all told, that’s six essays on Fontigens, written over a span of 16 years, with which I expect you to be familiar, before going forward into the blog post below. What, no? Tut-tut! See note [4] below for your homework assignment.
Moving forward now, cautiously. Down toward the bottom of my essay of [3July19] I wrote, “together Hsiu-Ping (Liu) and I worked up a small proposal to the Bernheim Board for a study on the evolution of Fontigens across the eastern USA.” I am here pleased to report that our proposal was funded just two weeks after that essay was posted, and that we subsequently sampled 13 additional populations of Fontigens beyond the singleton putative F. cryptica that kicked off our research effort, and that the results are startling, on a malacological scale, sort-of, I suppose.
 |
| Topotypic F. nickliniana (Font14) |
Hsiu-Ping and I settled on a study of sequence divergence in the mitochondrial CO1 gene quite early in our study. But as I have emphasized many times, indeed as many as the 18 posts currently indexed at the margin of this blog under “Gene trees,” molecular phylogenies are dependent variables, not independent variables. By themselves, they are at best weak, null models of population relationship. Only the scientist who brings with him a hypothesis regarding the evolution of a group of organisms can understand a gene tree that might be derived from that evolutionary process.
Thank heaven, then, that the 1990 monograph by Hershler, Holsinger, and Hubricht on the North American Fontigens has been sitting at the front of the thick Hershler file in my reprint cabinet for thirty years [5]. Last month [6July22] I sang the praises of that wonderful work – complete, scholarly, detailed, and unsullied by any taint of molecular phylogenetics. Entirely upon the basis of morphological data, demonstrating a remarkable appreciation for both intrapopulation and interpopulation variation, my buddy Bob recognized nine species of Fontigens in North America, plus F. cryptica of “uncertain status.” I cannot imagine any finer standard against which to calibrate a gene tree such as the one Hsiu-Ping and I proposed to construct.
So, the late summer of 2019 found me cruising along the Blue Ridge of Old Virginia, very much as I described in my essay of [26July06]. And as my idyllic journey unfolded, I paused to collect three of the many populations of Fontigens orolibas Bob Hershler catalogued in his monograph. These included Leslie Hubricht’s (1957) type locality at Hawksbill Spring (Font17), beside the Appalachian Trail at northbound mile 930.5, a prettier habitat it is difficult to imagine. And I sampled a second F. orolibas population (Font18) just 80 km south on the Blue Ridge Parkway at the pioneer springhouse figured in my 2006 post. And our good friends Wil Orndorff and Tom Malabad of the VaDCR contributed a third population of F. orolibas from Hugh Young Cave (Font19), 250 km further SW, to slide the scale.
And ditto for Fontigens nickliniana. Although Isaac Lea’s (1838) type locality for “Paludina” nickliniana (Hot Springs, Virginia) is obscure, I felt as though a sample from Blowing Springs (about 9 km N of Hot Springs) could reasonably substitute (Font14). Then I added a second population of F. nickliniana listed by Bob Hershler 140 km NE at Lantz Mills, VA, (Font15), and a third population about 250 km SW at Fleenor Spring, VA, (Font16). To these samples Hsiu-Ping was gratified to contribute an individual F. nickliniana she found in her freezer from Indiana (Font3). And you might remember, but are excused if you do not, that as of 2019, GenBank already held exactly one CO1 sequence for any Fontigens whatsoever, that of Fontigens nickliniana, an individual collected in Michigan [6].
 |
| Blowing Springs, VA [7] |
I was also able to sample the type locality of Fontigens morrisoni on my 2019 field trip through the green rolling hills of Old Virginia (Font13). Hsiu-Ping had samples in her freezer from the type localities of F. tartarea (Font1) and F. bottimeri (Font6), and our colleague Bob Weck sent us samples from the type locality of F. antroecetes in Illinois (Font22). And our good friends Wil and Tom contributed a second population of F. bottimeri sampled from Ogden’s Cave about 100 km W of the type locality (Font11). Comparison to the genetic diversity demonstrated by these 13 carefully-chosen populations (plus the singleton sequence from GenBank) should, we felt, allow us to place the CO1 sequence divergence demonstrated by our single putative F. cryptica individual from Kentucky (Font12) into its proper evolutionary context.
A neighbor-joining gene tree based on our CO1 results is shown below [8]. And the most striking result of our analysis was, without question, the tremendous mtDNA sequence divergence we discovered among conspecific populations. Sequence variation within the Blowing Springs type population of F. nickliniana (Font14) was negligible, for example, and ditto within Font16 from Fleenor Spring, and ditto within Font15 from Lantz Mills – just a few nucleotides. But in striking contrast, the mean between-population sequence divergence was 10.8% between Font14 and Font16, 10.4% between Font15 and Font16, and 11.0% between Font14 and Font15.
Similarly negligible values of sequence variation within populations, contrasted with strikingly-high sequence variation between conspecific populations, prevailed across our entire control sample of six Fontigens species. Mean interpopulation sequence divergence reached as high as 14.3% between the F. orolibas inhabiting the Hawksbill Spring type locality (Font17) and Hugh Young Cave (Font19).
This situation was so reminiscent of the population genetics characteristic of pleurocerid snails [9] that I suggested to Hsiu-Ping we might want to look for mitochondrial superheterogeneity within our three F. nickliniana populations. We upped our samples sizes to 30 for Font14, 10 for Font15 and 10 for Font16, but didn’t find any.
The levels of interpopulation mtDNA sequence divergence we discovered within species did not, however, swamp out divergence between the species. The mean CO1 divergence among species ranged from a high of 21.0% (between F. nickliniana and F. morrisoni of Virginia) to a low of 8.7% (between Virginia F. bottimeri and Illinois F. antroecetes).
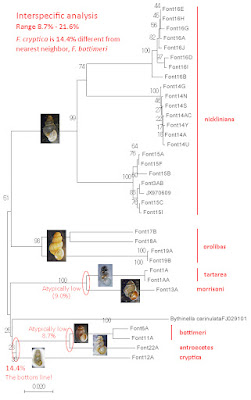 |
| NJ tree from CO1 sequence |
Our second-most striking result was the nesting of the two F. nickliniana sequences we analyzed from the Midwest, Font3 from Indiana and JX970609 from Michigan, within our Font15 cluster from Lantz Mills, Virginia. I admit that I do not have as much experience pioneering through forests of gene trees as most of my colleagues. But this is the clearest example of a “shared primitive” mtDNA sequence I have ever seen. Cladists made up the term “symplesiomorphy” to describe this phenomenon and used it for tails on fish.
And finally. The only unknown sample hanging on our entire tree was Lori Schroeder’s tiny snail, Font12, the singleton individual that she and Andrew Berry discovered under that rock at the springhead in the Bernheim Forest back in 2019. The minimum mean CO1 sequence divergence between that little snail and any of the control Fontigens species in our study was 14.4%, with F. bottimeri of Virginia. Thus, our hypothesis that Lori’s snail was sampled from a distinct species is confirmed. And since the shell and the animal both match Leslie Hubricht’s (1963) description of Fontigens cryptica [10], it is F. cryptica.
The confirmation that a population of Fontigens cryptica inhabits the subterranean waters of central Kentucky is not surprising, I don’t suppose, especially after five years of breathless advertisement in the columns of this blog. But the high levels of interpopulation mtDNA sequence divergence we documented across the genus Fontigens on the way to that unsurprising conclusion, my goodness!
Twenty years of published research on hydrobioid evolution in the American West led us to expect much less than 1% divergence between conspecific populations. Bob and Hsiu-Ping haven’t typically documented much more than a couple percent CO1 sequence difference among their scores of newly-described species. But in Virginia, even a very small sample of three F. orolibas populations returned interpopulation sequence divergences as high as 14.3%! Has the evolution of hydrobioid populations in isolated springs of the American West been so very different from the hydrobioids inhabiting isolated springs of the American East? Startling, indeed.
Notes
[1] Liu, H-P., L. Schroeder, A. Berry, and R.T. Dillon, Jr. (2021) High levels of mitochondrial DNA sequence divergence among isolated populations of Fontigens (Truncatelloidea: Emmericiidae) [2] in eastern USA. Journal of Molluscan Studies 87. [PDF] [html]
[2] No, the genus Fontigens has not properly been assigned to the family Emmericiidae since 2013 [3]. The parenthetical material was added to our title by some clueless naïf in the editorial office of the journal.
[3] Wilke T., Haase M., Hershler R., Liu H-P., Misof B., Ponder W. (2013) Pushing short DNA fragments to the limit: Phylogenetic relationships of “hydrobioid” gastropods (Caenogastropoda: Rissooidea). Molecular Phylogenetics and Evolution 66: 715 – 736. For a review, see:
- The Classification of the Hydrobioids [18Aug16]
[4] Here is your homework assignment. Yes, this will be on the test. All of it:
- Springsnails of the Blue Ridge [26July06]
- Lori Schroeder’s tiny snails [17July17]
- The most cryptic freshwater gastropod in the world [6Aug17]
- Not finding Fontigens cryptica [6Sept17]
- Finding Fontigens cryptica [3July19]
- My buddy Bob [6July22]
[5] Hershler, R., J.R. Holsinger & L. Hubricht (1990) A revision of the North American freshwater snail genus Fontigens (Prosobranchia: Hydrobiidae). Smithsonian Contributions to Zoology 509: 1-49.
[6] In my essay of [18Aug16] I mentioned that “a Fontigens nickliniana sample from Michigan” was the only fontigentine included in the gigantic worldwide CO1+16S+18S gene tree of Wilke and colleagues [3].
[7] Air exactly 58 degrees F blows out of pipes set in the mountainside above the spring, 365 days per year. I don’t know why.
[8] Lori showed this gene tree in a talk she gave at the OVUM meeting in 2019. A couple notes might be helpful for those of you comparing the 2019 tree to the one we published as Figure 2 in our paper [1]. We changed our population codes slightly between 2019 and 2021, so that Font14 became “nic14” and Font17 became “oro17” and so forth. And the figure in our published paper was a Baysian tree, rather than simple neighbor-joining. And we added a couple additional outgroups from GenBank.
[9] The phenomenon of double-digit mtDNA intrapopulation sequence variation is so common in pleurocerid populations that I have coined the term "mitochondrial superheterogeneity" to describe it. For a review, see:
- Mitochondrial superheterogeneity: What we know [15Mar16]
[10] Hubricht, L. (1963) New species of Hydrobiidae. Nautilus 76: 138 - 140.








So, you found F. cryptica . . . but did you find Bob?
ReplyDeleteNo, alas. The abundance of Fontigens cryptica seems now to have exceeded the abundance of Bob Hershler.
Delete